What is S-acylation?

S‑Acylation (often called palmitoylation) is a reversible lipid modification in which a fatty acid is attached to a cysteine residue of a protein via a thioester bond (hence the “S‑”).
A large number of diverse proteins are modified by S-acylation, and indeed the fatty acid chains added to these proteins are also diverse, and include palmitate (C16:0), stearate (C18:0) and oleate (C18:1). S-acylation can impact the trafficking, localisation, and stability of proteins and has emerging roles across several physiological systems and disease mechanisms.
Vision
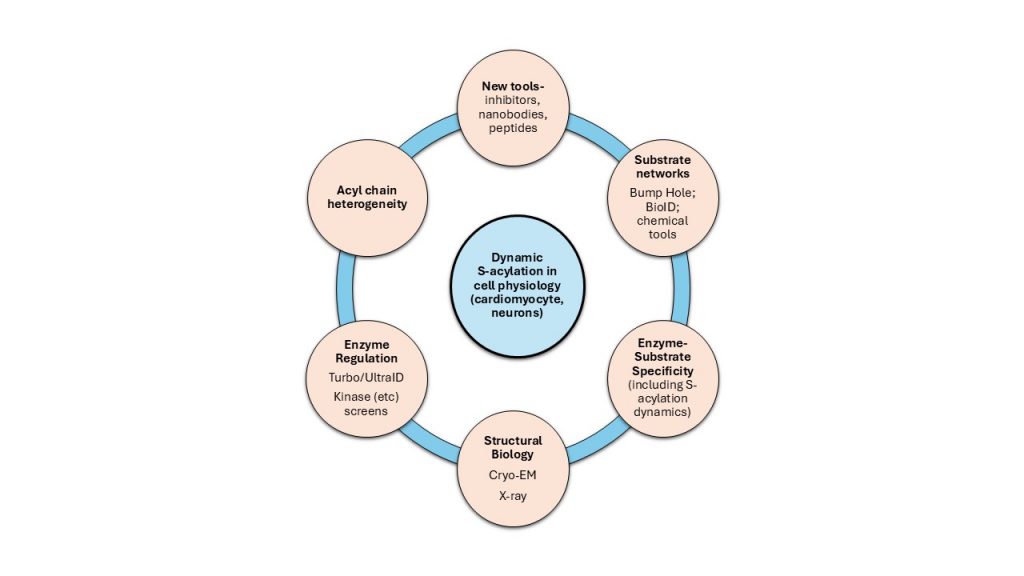
Our vision is to work as an integrated interdisciplinary team to deliver fundamental insights into the role of S-acylation in cell physiology. We will discover and design novel chemical modulators of S-acylation, map the substrate networks of individual acylation and deacylation enzymes, uncover structural details of enzyme-substrate recognition, identify novel mechanisms of enzyme regulation, and determine how acyl chain diversity affects S-acylated protein function. By integrating these discoveries, we aim to establish a comprehensive understanding of S-acylation’s role in cellular processes.
In addition to providing a major breakthrough in the fundamental understanding of S-acylation, this work aims to unlock its potential as a novel drug target with industrial, clinical, and societal impact.
The Collaboration
Our collaboration brings together teams of experts from across the UK, located at Coventry University, Imperial College London, the University of Glasgow, the University of Oxford, and the University of Strathclyde. We are an interdisciplinary team with expertise in biochemistry, cell biology, chemical biology and structural biology. The teams provide expertise in the following areas:

Beveridge: expertise in the use of native and intact protein mass spectrometry. This expertise will be applied to measure the stoichiometry of S-acylation, as well as detect heterogeneity in the lengths of the acyl chain modifications

Chamberlain: expertise in analysing enzyme-substrate interactions to elucidate how zDHHCs selectively recognise and modify target proteins within their specific substrate networks. This expertise will be applied to study substrate recognition by the zDHHC enzymes that are the focus of the sLoLa programme, and to contribute to the mapping of zDHHC substrate networks.

Fuller: Expertise in quantitative, mechanistic analysis of S‑acylation and how it controls cardiac ion channels, transporters and signalling molecules, with emphasis on how S‑acylation modulates excitability, calcium handling, and cardiac performance. This expertise will be applied to define how selected zDHHC enzymes regulate electrical and biochemical signalling in cardiomyocytes, and to integrate these molecular mechanisms with functional readouts to map enzyme-substrate relationships within the heart.

Greaves: Expertise in analysing the molecular mechanisms by which deacylase enzymes regulate dynamic S-acylation, including how these enzymes selectively recognise and interact with individual substrates and how their enzymatic activity is controlled. This expertise will be applied to investigate substrate recognition by the deacylase enzymes that are the focus of the sLoLa programme, and to contribute to mapping deacylase substrate networks.

Siebold: Expertise in Cryo-Electron Microscopy and tomography, X-ray crystallography, imaging, and protein-protein interactions, which will be applied to the structural characterisation of zDHHC enzymes and their substrate interactions

Tate: Expertise in high-throughput chemical biology, drug discovery, proteomics, and functional genomics. This expertise will be applied to map the substrate networks of zDHHC and APT enzymes and to develop novel binders and chemical modulators of these enzymes that will support their structural and functional characterisation.

Tomkinson: expertise inDesign, synthesis and biological evaluation of chemical tools to develop an understanding of enzyme-substrate networks, enzyme-substrate selectivity and substrate-acyl chain location/identity. In addition, we will develop chemical probes to inhibit the zDHHC family of enzymes to allow the dissection and interrogation of their biological mechanisms and pathways. analysing enzyme-substrate interactions to elucidate how zDHHCs selectively recognise and modify target proteins within their specific substrate networks

Walden: Expertise in structural biology methods (X-ray crystallography and cryo-electron microscopy) and biophysical techniques, which will be used in structure determination and assay development for understanding how deacylating enzymes recognise and bind their substrates, as well as elucidating potential substrate selectivity.
Strategic Advisory Board
The project is overseen and guided by a team of International experts in related fields. Members of the SAB include:
- Maureen Linder (Cornell University)
- Rami Hannoush (Versant Ventures, CA)
- Bryan Dickinson (University of Chicago)
- Ari Sadanandom (University of Durham)
- Virginia Cesare (University of Dundee)
Funding

The project is funded by the BBSRC Strategic Longer and Larger (sLoLa) programme.
“An Integrated analysis of S-acylation dynamics and its importance in cell physiology” BB/Z517409/1
